Second Takes
Andrea Thorn
University of Hamburg
Published February 28, 2023
Andrea Thorn had been a junior group leader for a less than a year when, in early January 2020, researchers in China identified the cause of a mysterious contagious illness with pneumonia-like symptoms. Less than a week later, they released its genetic code, which revealed the novel virus to be a close cousin to the SARS coronavirus (SARS-CoV-1) that caused an outbreak in 2002-2003.
The rapid scientific response gave Thorn an idea for a talk she was due to give over coffee and cake to colleagues in her building at the Julius Maximilian University of Würzburg in Germany. She ran the only lab there focused on new computational methods for experimental structural biology. Most scientists in the building were infectious disease specialists.
"I needed to explain to them how my work was relevant," she says. Thorn works on ways to better model the atomic structures of RNA and proteins to make them useful for structure-based drug discovery, something that quickly would become more important as researchers rushed to respond to COVID-19.
Molecular models are a foundation of modern drug and vaccine development. They enable scientists to identify how to break the cycle of infection by targeting the right spot in the right protein. But even the most carefully determined structures reported in the most prestigious journals are imperfect interpretations of experimental data, Thorn says. Small errors can have large consequences in the search for a drug with the right fit.
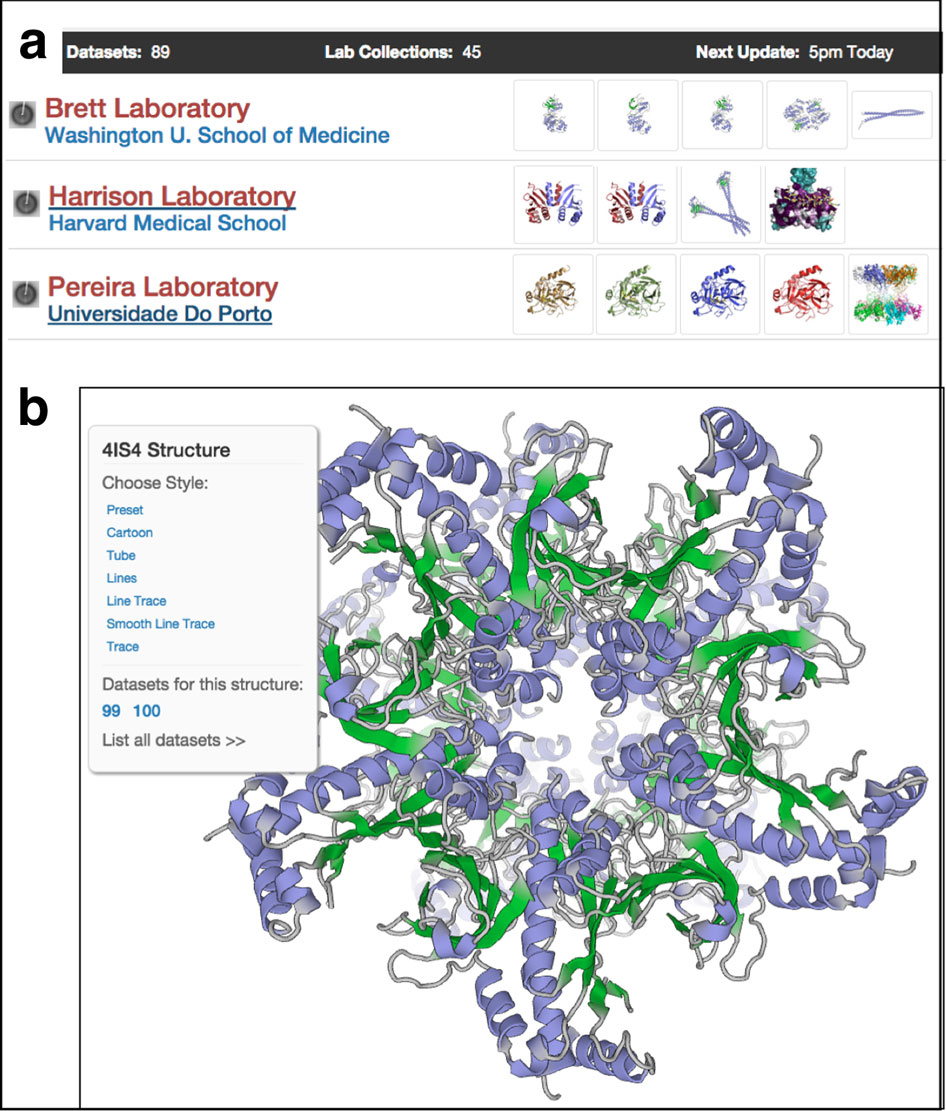
When Thorn was preparing for her talk, no structures from the SARS-CoV-2 virus had been published yet. But more than 100 structures from SARS-CoV-1 were available in the worldwide Protein Data Bank (wwPDB). The structures ranged from a key protease in viral replication to the spike protein needed to attach and enter host cells.
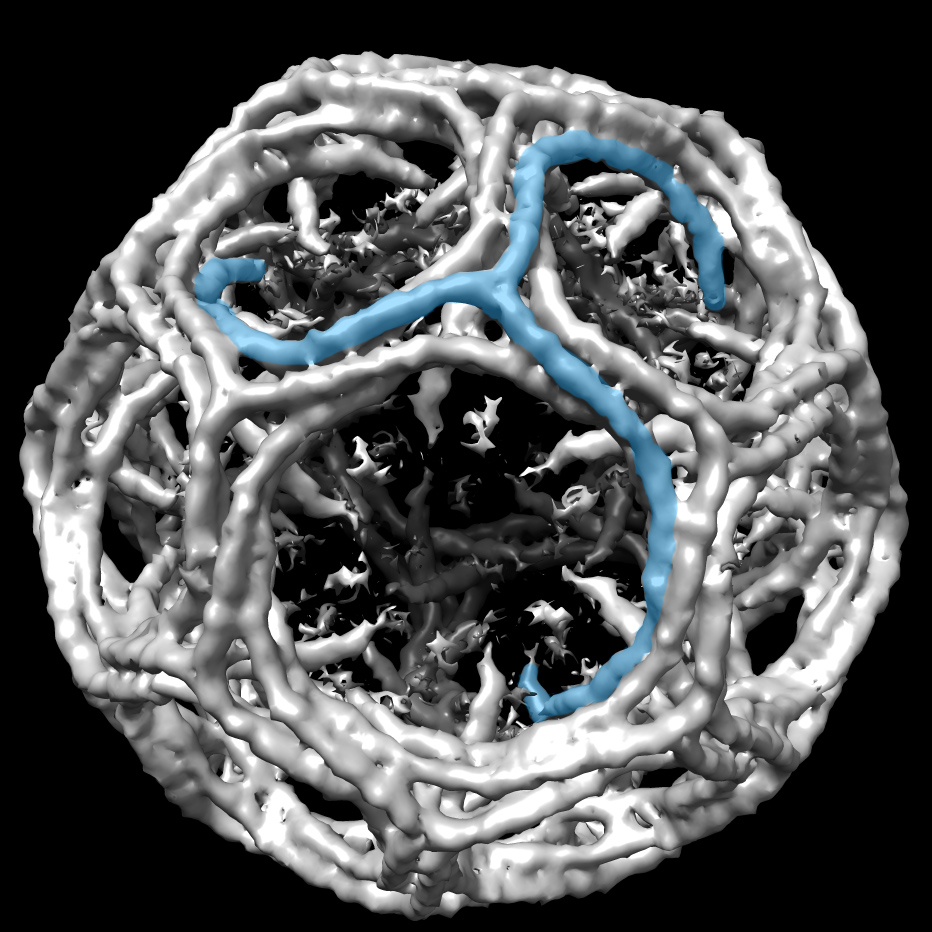
From a random sample of five, Thorn found three SARS structures to illustrate the point of her research: Most protein structures can be improved with additional analysis.
Her talk was well received, germinating the seed of another idea that first seemed outlandishly bold and then became increasingly urgent as COVID-19 spread and the death toll soared. Thorn floated the idea over dinner with her mentor, Arwen Pearson in Hamburg: Analyze and correct all SARS structures. Pearson encouraged her, as did another senior colleague, Elspeth Garman of Oxford University.
In March 2020, as the first SARS-CoV-2 structures were being reported, Thorn first got her own small group on board and within a week pulled together a team of like-minded and mostly junior structural biology developers from all over the globe as the “Coronavirus Structural Task Force”. Many of them were international specialists in their respective fields, she notes.
They met online every weekday and posted daily updates to GitHub for anyone to access. Every Wednesday, when new wwPDB structures were released, their automated pipeline identified new coronavirus structures and assessed the quality of a representative sample of models and experimental data. When they improved a structure, they sent it back to the original authors to update the wwPDB entry, no strings attached.
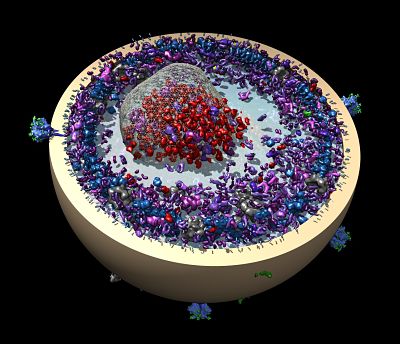
For their web site (https://insidecorona.net), the group wrote blog posts to share with colleagues the larger story emerging from aggregated data of multiple structures. They wrote explanatory pieces for the public. They posted a 3D printable model of SARS-CoV-2.

A year into the pandemic, structural biologists had released a total of 1,146 SARS structures covering 18 proteins from both SARS-CoV-1 and SARS-CoV-2. Most were derived by X-ray crystallography (73%) and single particle cryo-electron microscopy (cryo-EM) (24%) (Nature Structural & Molecular Biology, January 2021).
The task force's analytical pipeline deployed software programs from contributors inside and outside the Task Force. Among them were two tools previously developed by Thorn—AUSPEX, used for experimental X-ray data to detect sample measurement and processing problems (Acta Crystallographica, 2017) and, for experimental cryo-EM data, HARUSPEX, a neural network tool to automatically distinguish between nucleic acids and protein and to assign secondary protein structure elements (Angewandte Chemie, International Edition, 2020).
"We didn't do any peer reviewed publication until 2021," Thorn says about the task force. "We just wanted to fight the pandemic. The whole thing was fast. We pushed and pushed and pushed against corona so that drug developers and vaccine developers would have the right data. You know, it's been fantastic. It was born out of spontaneous willingness. I wish science would always be like this."
The task force published a description of its work (Nature Structural & Molecular Biology, May 2021). In 2023, Thorn says a dozen articles are in progress for peer-reviewed journals to summarize the combined information and results of their work, as well as to note questions yet to be answered about SARS-CoV-2. The task force will wind down in summer 2023. Thorn is strategizing on the next steps for a tenured academic position.
Thorn grew up in Germany in an intellectual family. Her father is a chemist, her mother an artist and entrepreneur. She is the third-born child with three brothers.
She calls her path to structural biology straightforward. One of her early science-related memories goes back to age 3, when her father explained surface tension to her with the gravity-defying demonstration of floating a needle on water. "My dad showed that to me with his always huge confidence that I would understand everything," Thorn says.
The family moved four times when Thorn was in elementary school. She supplemented the patchwork early education with natural curiosity. She read voraciously from the family library, made observations with a home microscope, and conducted chemistry experiments on the sly. In her mother's atelier, she also painted on large blank canvases.
In her teens, she and her friends were early adopters of live action role-playing games that have subsequently become popular in Germany. Participants assume the roles of characters and act out scenarios in a collaborative storytelling experience. To prepare for her roles, she researched scenarios ranging from futuristic military campaigns in Libya to glial cell retrieval from patients. "I read up to 600 pages a day when I was a teenager," she says.
In school, Thorn's studies became concentrated in chemistry and art. She continued first to a bachelor's degree and a master's degree, a prerequisite for her PhD.
She excelled in the lab and in computational chemistry, but her breakthrough lessons came after she failed quantum chemistry. She sought a tutor in the solid-state physics department, where her academic track record was unknown. A professor she consulted for a referral, Helmuth Zimmermann, volunteered to oversee weekly tutoring for Thorn and other classmates who needed help.
"Arguably those tutoring sessions did more for my education than everything else that semester," she says. "I'd never used maths before to describe things in nature in that way and fell in love with that."
One day, in response to Thorn's curiosity, the professor explained his research, which was measuring crystals to determine the structure of molecules. "I was hooked, because that meant the maths that I just learned could be applied far beyond class," she says. "It meant you could use them to find out the structure of crystal lattices and, with the structure of crystal lattices, the structure of molecules."
She skipped her prescribed medicinal chemistry classes in favor of crystallography lectures, which were not in her curriculum plan. One day, her crystallography lecturer refused to answer her question about SHELX, a crystal structure refinement program, saying she wasn't smart enough to understand.
"I was so angry, I printed out the SHELX manual, which is 100 pages, and I read the whole fricking thing." She came to the next class armed with new and more detailed questions. "It turned out he didn't know so much about crystallography," she says.
When it came time to choose a PhD group, Thorn had a list. The list included George Sheldrick at University of Göttingen, who developed SHELX. On the way to a holiday live role-play destination, she stopped by his lab to scout it out informally. Unexpectedly, she had a long conversation with him that ended with an immediate offer of doing her PhD thesis in his group.
She thrived in the high-performing lab and found fellow role-play enthusiasts. When she was teaching her first classes as a PhD student, a friend came to collect her for lunch. He noted her stance in front of the students (who were much older than her) looked like the last weekend's live role play when she had played a commanding leader with feet planted and shoulders squared as she led her warriors into battle.
In 2011, Thorn graduated with her doctorate and four new first-author publications from her thesis. The morning after her defense, she married her childhood sweetheart, a chemist. Within a year, she had secured a prestigious three-year Marie Curie fellowship for independent research at the MRC Laboratory of Molecular Biology in Cambridge, followed by a brief but productive stint at University of Oxford.
Homesick for Germany and their parents, she returned with high expectations of becoming an assistant professor, something that remains a goal. She secured outside funding and became a group leader at University of Würzburg in 2019 and moved to Universität Hamburg in 2020.
The task force may be winding down, but its value lingers in Thorn's mind as an unmet need in science. She has been thinking about the volume of data being generated by the structural biology community and how to combine the data for more meaningful interpretation and usefulness. She also takes it as an example of what is possible when experts across the globe start to collaborate on an important problem.
"We are generating data like crazy," she says, “beyond the capacity of individuals to assimilate and make sense of details from different techniques across labs. "In Germany, we have this beautiful word, Datennachnutzung. It means you are using experimental data for a second time to find new insights. And that's something we absolutely need to do."
-Carol Cruzan Morton