The Fixer
Jane Richardson
Duke University
Published October 28, 2013
In the late 1960s, only a dozen or so proteins had been solved using x-ray crystallography. Jane Richardson and her husband, David, solved one of them (Staphylococcal nuclease), while working at MIT and a second of the first 20 (superoxide dismutase) at Duke University, where they still work today. The problem was, even with the solutions in hand, no one could quite comprehend all the complex information in such structures. There was no standard way of visualizing them.
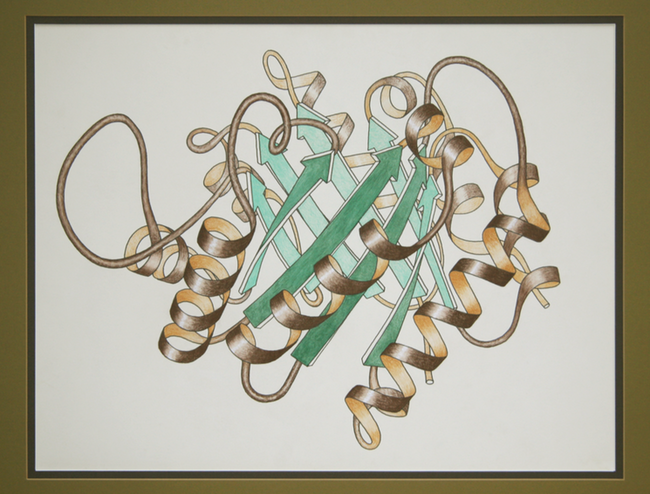
So Richardson, now a James B. Duke professor of biochemistry at Duke University, spent two years teasing out a method for drawing and labeling these structures. Her ribbon drawings and seminal 1981 paper, "The Anatomy and Taxonomy of Protein Structure," became the standard for visualizing proteins. "It's probably the most notable thing I've done in my whole career," says Richardson, who without formal doctoral training has advanced in academia, earning prestigious awards such as a MacArthur Fellowship in 1985, and election into the National Academy of Sciences, the American Academy of Arts and Sciences, and the Institute of Medicine.
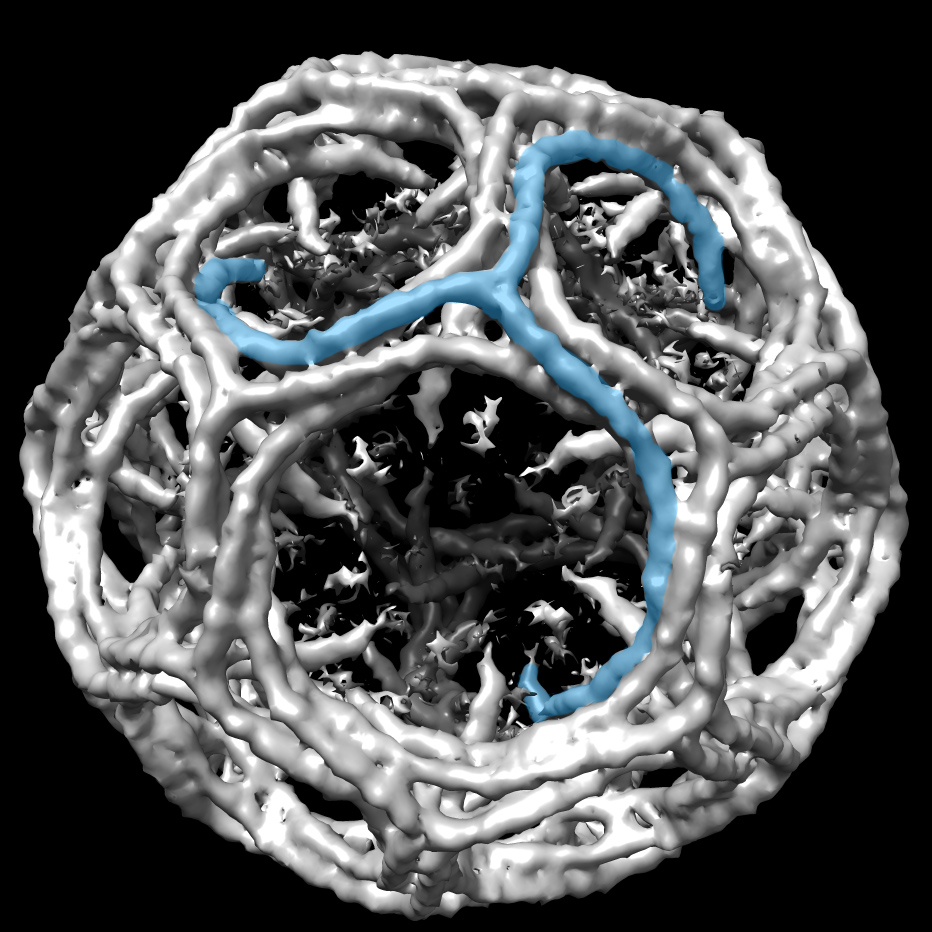
Working before the rise of computer graphics programs, Richardson hand-drew 75 different structures. In doing this painstaking work, she developed an intuition about proteins that marks, essentially, the very beginnings of structure validation. Richardson could just tell when a solution wasn't right. "It's having looked at the detailed structures," she says. "Some of it you can get at the schematic level, where you know they haven't done it quite right. The most obvious one is if there isn't much of any secondary structure. People always under assign secondary structure at low resolution."

Since then structure validation and the necessary partner process of repair have become the primary focuses of her career.
From Pen to MolProbity
In the early 1980s, Richardson was a non-tenured "Associate" in the lab of her husband, David Richardson, professor of biochemistry at Duke University. Their joint research was at the forefront of synthetic and computational biology, doing de novo protein design. Their aim was to design for structure, rather than function. According to Richardson, they had some success, in usually getting the right overall folds. But they never got well-ordered unique proteins. "We got molten globules," she says.
They realized that they weren't getting the internal contacts right. "You really can't think about molecular contacts either inside a structure or between molecules without having all the hydrogen atoms there. They're what make all the contacts," she says.
By necessity, the Richardsons began looking into adding the hydrogens. This began what would become a 5-year effort to develop methods to add the hydrogens to a structural model and look at their contacts. They named the tool ALL-ATOM CONTACTS. This and other related tools, such as REDUCE, which optimizes the hydrogen contacts, and PROBE, which shows them graphically, gave shape to the larger software package now called MolProbity.
Before MolProbity, validation tools developed in other labs, such as PROCHECK, first available in 1993, and WHATCHECK, developed in 1997, provided statistical evaluations of solutions, applying Ramachandran and rotamer distributions, for example, to flag problem areas.
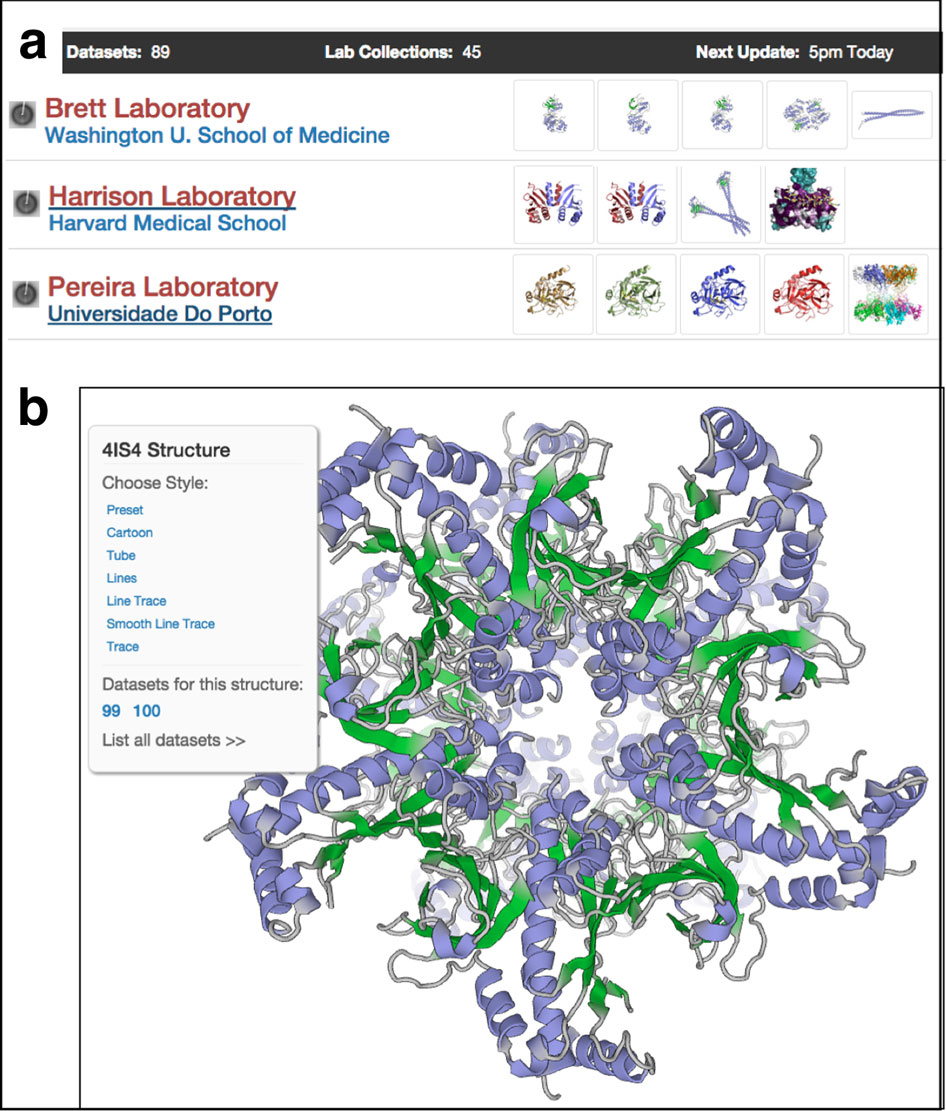
Today, MolProbity has also implemented these statistical methods for validation. The Richardsons' original implementation of rotamers in MolProbity relied on a database of the top 100 structures. They now are working on an update that uses an 8000-structure database.
The additional data makes validation more complex. "Almost nothing is actually a normal distribution," she says. "Most of the errors are systematic errors rather than random errors. As a programmer, you have to try to understand what causes the systematic error, so that you can do a much better job of fixing it."
The validation parameters used by MolProbity come from a combination of a small molecule database and from the well-ordered parts of the high-resolution Protein Data Bank (PDB) structures. Other validation parameters used by MolProbity come from a landmark paper written in 1992 by Engh and Huber that defined bond lengths and angle parameters for proteins. "Everything is still pretty close to those classic parameters," says Richardson. "We've recently spent a couple of years sweating over a redo of the hydrogen parameters and we've almost finished it. We've gotten them a whole lot better than they were."
ABCs of PDB Validation
The PDB is a vast database of protein structures. In recent years, it has put structure validation front and center, using tools like MolProbity to score submissions, providing a means for users of the database to assess the quality of a structure. Richardson is a member of several validation committees for the PDB. She helped write the validation requirements for x-ray crystallography that came out in 2011.
From Richardson's point of view, validation should be a way of life for the crystallographer, not an afterthought. "People want to leave things open because, for instance, you can get rotamer outliers that are real, even occasionally Ramachandran outliers that are real," says Richardson. That, however, is the exception and should not be counted on. "The best bet initially is to set things in the expected places, then see what the data is telling you. If the outlier is real, it will win out. Doing it the other way around is very dangerous."
Richardson also has several outstanding concerns about structure deposition. One is regarding the fact that the PDB does not currently allow hydrogen atom coordinates in a deposited file. "Some people think that it's trivial to add them and that everyone would add them the same way. That is simply not true," she says.
Another concern is about resolution, an important quality measure in protein crystallography. The problem is, the measure is not actually well defined. Data sets are too asymmetrical and peculiar and inconsistent from one experiment to the next. Consequently, depositor decisions about how to define resolution are somewhat subjective and arbitrary. "From our point of view it is a real nuisance," says Richardson. "If you look from 10,000 feet, resolution is perfect as the independent variable. It is the basic measure of how much information you have, going in. But the detailed view isn't great."
There are many competing proposals, but little agreement on which one should win out. "We'd love to have a standard definition even if it isn't perfect," she says.
The Devil in the Details
At the present, MolProbity does not validate data, nor does it validate how well the model matches to the data. However, MolProbity is currently undergoing a major renovation. A research associate in the Richardsons' lab, Jeff Headd, is rewriting the underpinnings of MolProbity in Python so that it will be compatible with Phenix tools. The goal is to build a more effective connection with Phenix and its crystallography-based tools. This will allow MolProbity to pull up refinements and maps and crystal symmetries and enable the addition of new validation features, says Richardson.
Another focus of their work is generalizing the methods for validation and repair to work better at low resolution. It's hard, she says, because there isn't enough data. "But we're trying to put in more of the info from the combined knowledge of 100,000 structures and try to do a better job with it," she says.
Her lab is also working on adding features to the ROSETTA and ERRASER programs, both developed in other labs, to allow ERRASER to do RNA structure corrections. "It's just amazing how different DNA and RNA are, given that the only fundamental difference between them is an extra OH-group on the sugar ring. That makes an enormous difference to how they interact," she says.
While DNA wants to form a helix structure to fulfill its primary and predominant function as an information storage medium, RNA has many different forms. "RNA is more like proteins in terms of overall tertiary structure and the fact that it can do catalysis and specific binding. It has many more conformational states than DNA," she says.
Oddly, says Richardson, it's easier to deal with RNA in validation than DNA because RNA conformations are "sort of rotameric." Sort of. As with the many other validation problems Richardson has worked on over the years, the devil is in the details.
-- Elizabeth Dougherty
Jane Richardson photo by Rita Lo