U-Store-It
The SBGrid Data Bank provides an affordable and sustainable way to preserve and share structural biology data
Published March 28, 2016
Evidence of the Higgs boson appears as a bump on a histogram resulting from the analysis of data from millions of detectors at the Large Hadron Collider. What if all that raw data vanished, leaving nothing but the histogram? The physics community would reel.
Yet in structural biology, raw data frequently goes missing. Scientists dutifully store models of proteins in the Protein Data Bank, but the X-ray diffraction data used to derive those macromolecular structures isn’t accessible easily, if at all. One reason is that, until recently, there was no clear place to put it.
Now, however, structural biologists can publish their data in the SBGrid Databank (SBDB data.sbgrid.org), a system designed to preserve and disseminate primary experimental datasets that support scientific publications. More than a repository, the system is a databank that is integrated into the SBGrid software distribution so that datasets are immediately available to anyone in the community. The SBDB is currently a pilot grid, but it will mature into a sustainable system over the coming three years with support from the Helmsley Trust Biomedical Research Infrastructure. This project is a collaboration between SBGrid, Dataverse, and Globus and the National Data Service has selected it as a pilot activity of.
“Diffraction datasets are often misplaced,” says SBGrid Founder and Director Piotr Sliz. “Those datasets are captured at state-of-the-art synchrotron facilities and are the ultimate outcome of long-term biological experiments. Publication of this primary data will support reproducibility, model validation, training and methods development.”
The Making of a Databank
The de facto approach to data management for structural biology is, with a few exceptions, every lab to itself. With terabyte drives selling for about $50 on Amazon, storage is cheap, so why not? The fact is, the simplicity stops there. Beyond storage, data needs to be archived, curated, and validated. To be valuable to the wider scientific community, it also needs to be shared. “The cost for storing data is small compared to the cost of validating it to make sure it is all ok,” says methods developer Tom Terwilliger, a structural biologist at Los Alamos National Laboratory.
The Protein Data Bank is the go-to repository for storing models, so it might also seem a natural place to store raw data. But the storage of diffraction data presents distinct challenges. “You have to worry about volume and distribution and also about making the data available to the community in a useful form,” says Sliz.
In 2015, in response to input from the structural biology community, SBGrid began piloting the SBDB to address these challenges. SBGrid has a long history of data management that stretches back to 2002, when it started managing diffraction data for structural biology laboratories at Harvard and affiliated institutions. In 2012, the Consortium also prototyped a system to move diffraction data sets between Harvard and two synchrotrons. The work created a foundation of experience with data annotation, replication, and preservation at SBGrid. “We’ve been worrying about data for quite a while,” says Sliz.
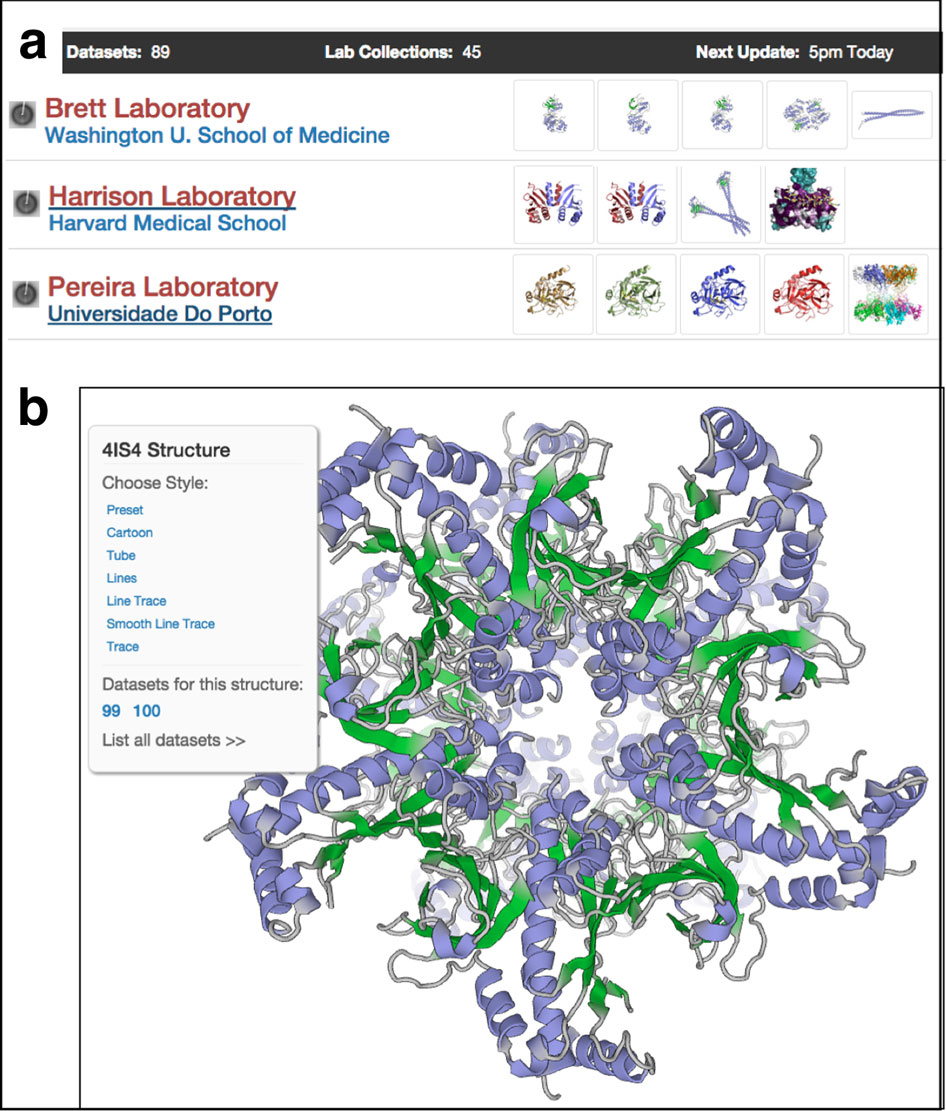
In this pilot data bank, data can be published quickly and accessed by SBGrid members as if it were local. Demonstrations of this capability are up and running at Yale University and Harvard University. “The datasets are part of the SBGrid computing environment,” says Sliz. “In contrast to a website repository, SBDB provides direct access to datasets from within computing environments.”
Sustainable Bytes
In the next phase of development, SBGrid is working to move the pilot implementation to an SBDB platform that can be managed, maintained and upgraded without incurring crippling costs. “As a community, we need to worry about the costs of supporting research infrastructures, and aim to preserve precious funding resources for experimental research,” says Sliz.
To accomplish this goal, SBGrid partnered with Mercè Crosas, the Director of Data Science at the Harvard Institute of Qualitative Social Science. Crosas and her team have over a decade of experience developing the Dataverse project, a fully featured repository for sharing large datasets in the social sciences. “The collaboration is good because it brings together two strengths: subject expertise about structural biology data from SBGrid, and from our perspective, expertise in data management,” says Crosas.

Mercè Crossas, Chief Data Science and Technology Officer at the Institute for Quantitative Social Science (IQSS) at Harvard University and co-PI for the Dataverse project for data sharing and archiving. Learn more about Dataverse project from their website dataverse.org.
The Dataverse framework is an open source solution backed by a community of developers who are supporting and expanding the features. It follows best practices for data sharing and publishing to ensure that data is discoverable and able to be referenced through citations both to and from journal articles and, in the case of structural biology, the PDB. “We need to make sure that long term access to the data is sustainable over time,” says Crosas.
In addition to sustainability, the SBDB will support a publication workflow that makes it possible for scientists to self-publish their data in the SBDB when they submit a paper to a journal and a model to the PDB. All three will be linked together through citations.
The planned workflow streamlines the data publication process by removing the initial gate of data validation prior to submission. Instead, SBGrid develops tools to complete post-deposition analysis and hopes to engage members of the community in the process.
Currently, SBDB automatically processes all X-ray datasets using XIA2, an expert data processing system. Dataset pages present the results. Some challenging datasets, however, might require manual processing and unique expertise for validation. “We want the community to be able to access all primary data as soon as possible and in parallel drive automatic evaluation to eventually catch up with cases that require more expertise,” says Sliz.
Currently, 84 percent of data sets in the pilot SBDB can be reprocessed to yield statistics similar to what was reported in corresponding publications. “That’s encouraging,” says Sliz.
This phase of the SBDB project, which is supported by funding from the Helmsley Charitable Trust, will take about three years.
Ultimately, what the SBDB is providing the community is not storage or data management, but a platform for doing science as it was meant to be done. “The SBDB makes it possible to re-determine any structure from the beginning with ever-improving tools, leading to ever-improving structures in the PDB,” says Terwilliger.
As SBDB becomes established, the SBGrid team is already thinking about expansion beyond X-ray diffraction data. Indeed, the data collection already includes microED, lattice light-sheet microscopy and molecular dynamics datasets. “This platform is applicable to other data-heavy fields of biomedical research,” says Sliz. “The same mechanism could be used to distribute other large datasets across the scientific community.”
-- Elizabeth Dougherty