Into Alignment
Geoff Barton
Division of Computational Biology, University of Dundee
Published May 27, 2015
In 1987, when Geoff Barton was a graduate student learning computational structural biology at the University of London, just 6000 protein sequences were known, but their numbers were rising exponentially, and it was becoming clear that they had commonalities. Sequences that yield valuable functions have staying power, so they are conserved throughout evolution. Finding these recurring patterns, however, required painstaking pencil and paper comparisons.

So Barton wrote a computer algorithm to align multiple sequences, an early contribution to what would become a prolific career in methods development. Today, Barton is Head of the Division of Computational Biology at the University of Dundee. About half of his lab develops methods and software of value to structural biologists. The other half focuses on genomics work, both looking at human disease and plant-based RNA. His scientific questions often drive his methods, which in turn, drive new hypotheses and collaborations. “I like to make things because they are useful to me, but I get a big kick out of making something that other people can use in ways I wouldn’t have thought,” he says.
The most well known of Barton’s programs is Jalview, which in more than one way was influenced by that early algorithm. For instance, Jalview is freely available. Barton’s first algorithm was not free due in part to the political climate in the United Kingdom at the time, which encouraged scientists to commercialize their work. As a result, his program was used, but when a similar but open access program called CLUSTAL came out, people flocked to it. “My program has about 500 citations. Not bad,” says Barton. “CLUSTAL has 60,000. It’s one of the most highly cited papers in any field.”Barton laughs about this now, be that experience taught him important lessons about the development of software for science. “Make it easy to use,” he says. “And give it away for free.”
Jalview development began in the mid-1990s at the European Bioinformatics Institute to provide an interactive way to align multiple sequences. Barton and lab member Michele Clamp, who is now at Harvard University, chose to write the program in Java, which was a brand new language at the time. The decision proved wise. Not only was the system well written and easy to use, Java made it portable and usable on the web, which was gaining in popularity at the time.
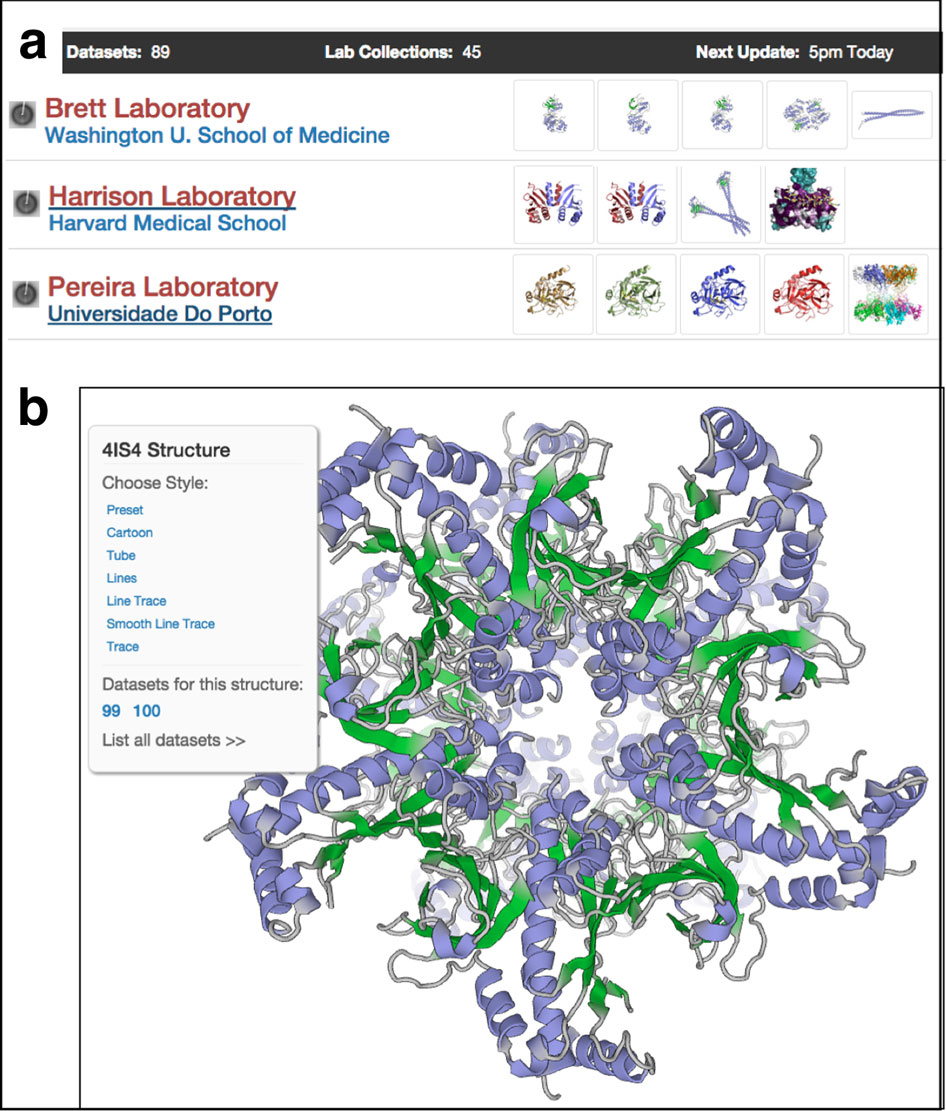
Barton calls Jalview “net centric” because, in a later version, his team added integration with multiple databases, including the Protein Data Bank, to bring in structural and annotation information, and multiple analysis services, including his own Jpred for structure prediction. JalView also farms out complex jobs to a computing cluster. Barton hosts a dedicated cluster at Dundee but users can specify a local cluster, too.
Very recently, Jalview added integration with Chimera. “Now users have the full power of a proper molecular graphics environment and Jalview together,” Barton says.
Currently in development is a feature that allows researchers to deeply explore sequence variations, such as human genetic mutations associated with a disease, in the context of other similar sequences. The goal is to help researchers gain insights about how different variations might alter structure and function. “I think it’s going to be powerful,” says Barton.
Barton’s own work is driving the development of this capability. His collaborators, a team of experts in the genetics of skin disease, found interesting and unexpected variations in whole exome sequencing data from patients with eczema. Barton hopes his variation analysis tools will help them narrow down the leads to guide future research and, eventually, drug discovery. “Often what bioinformatics is about is analyzing large data sets to prioritize what you should look at next,” he says.
Because of Jalview’s potential to accelerate research that could have major impacts on health, Barton and lab member Jim Procter were finalists for a Biotechnology and Biological Sciences Research Council (BBSRC) Innovator of the Year award in March 2015. The winning innovation, a diet that makes cows produce low-fat milk, was a tough contender. “We had a harder time explaining why Jalview was exciting,” says Barton.
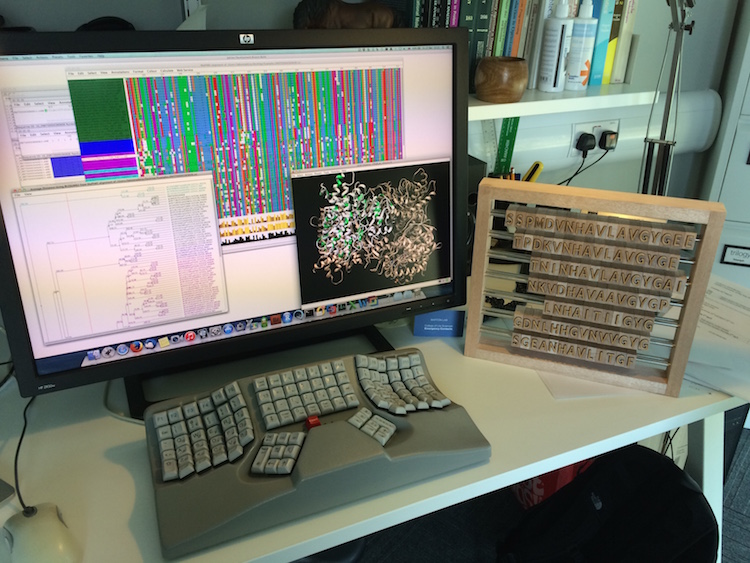
Other programs of particular interest to structural biologists coming from the Barton lab are JPred, which was recently updated, and TARO, a predictive tool that rates the likelihood that given protein sequence will be successfully overexpressed, purified, crystallized and solved. While Jalview is part of the SBGrid installation, many of Barton’s software tools are web-based, so they are not included in the distribution. Please visit his website (http://www.compbio.dundee.ac.uk/software.html) for a complete list.
-- Elizabeth Dougherty