Frame by frame
Nikolaus Grigorieff
HHMI’s Janelia Research Campus
Published January 28, 2019
Rejection is an unavoidable part of science, whether it’s funding for research or manuscripts submitted for publication. But few people showcase their setbacks like Nikolaus “Niko” Grigorieff, whose lab web site features a separate “rejection” section.
As a methods developer in structural biology, Grigorieff may be best known for pioneering a technique known as motion correction in cryo-electron microscopy (cryo-EM). But the numbers tell a different story.
All together, his papers have been cited about 9,500 times, excluding self-citations. Compare that to the 45,000 downloads of rejection letters his web site logged as of January 2019 for the 20 papers (so far) that were vetoed by one or more journals before being published.
“People love reading these rejections,” says Grigorieff, who started posting them in 2010 after lab discussions of frustration with the peer review process.
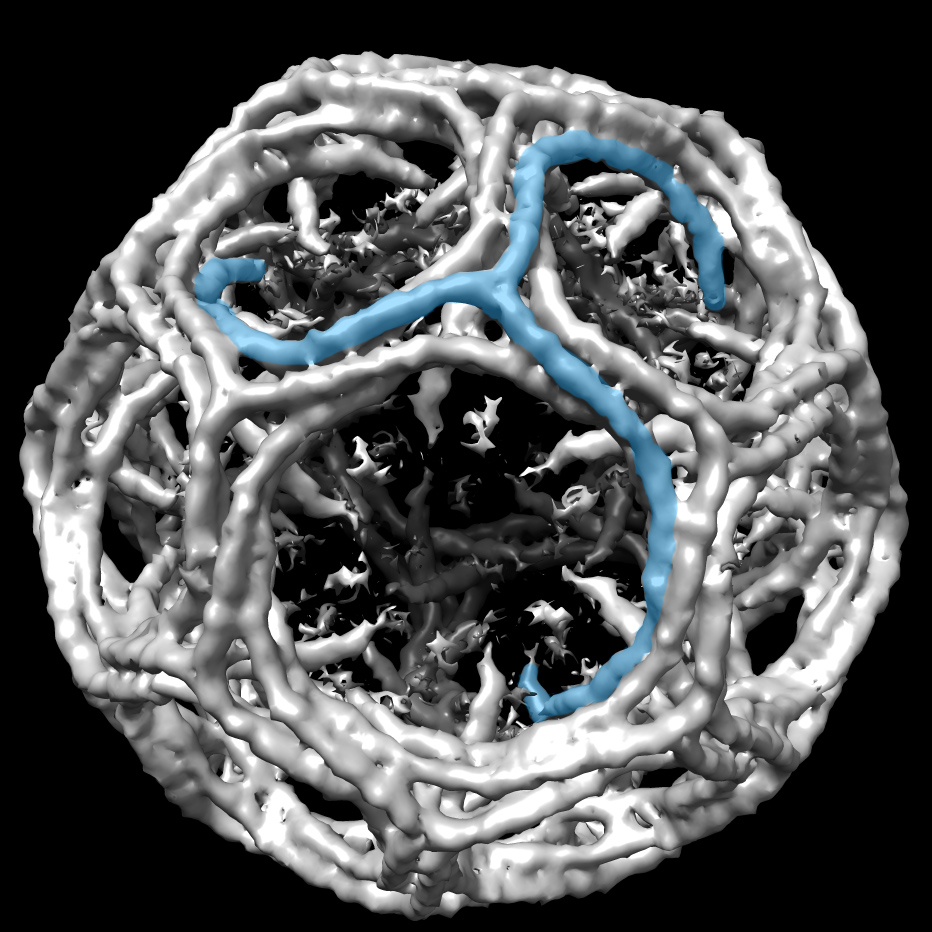
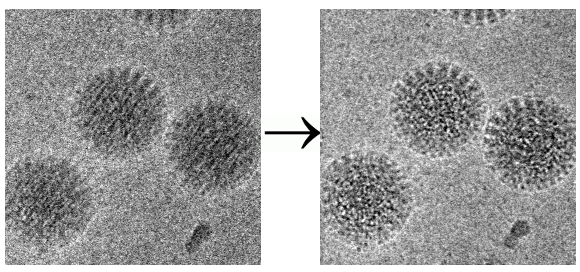
One notable rejection involves one of the group’s pivotal motion correction studies. It demonstrated how taking movies and analyzing them frame by frame can restore atomic detail to blurred molecular images. The rejection letters from Nature Methods and PNAS are posted with the PDF of the 2012 paper in Structure. “After this paper was published, everybody in the cryo-EM field who has a movie-capable camera switched to movies,” he comments on the entry.
Other authors are welcome to contribute their rejection letters as soon as a paper is accepted elsewhere, but so far only one other research lab has volunteered a sample.
Grigorieff grew up in Berlin, Germany. He didn’t like to hang out with people, preferring to fiddle with electronics and computers at home. His fix-it skills made him popular with friends and family, at least most of the time. After his mom turned off the electricity because he was playing his music too loud, he built a generator to power his stereo independently.
He even wrote his own computer games, including a strategic game that beat him a few times. These days, online gaming helps him maintain family ties. Since he moved away from Germany after his undergraduate degree, he has a Skype chat with his brother almost every Sunday, and then they play an online game together.
Grigorieff picked up his EM skills during his physics PhD training at the University of Bristol in the UK. For postdoctoral work, he moved to the Cambridge lab of Richard Henderson, where he encountered proteins and the related cryo-EM motion problem.
In materials science, “you switch on the electron beam, the sample starts moving, and you wait until it stops to take a picture. There, you can get high resolution without movies,” Grigorieff says. “But with a protein or a cell, it’s getting more and more damaged while you wait. By the time it stops moving, there's nothing left. You have to switch on the beam and take a picture immediately when the sample is still intact. But you had to accept that it was always a little blurred because of this movement.”
At first, Grigorieff helped develop the two-dimensional electron crystallography in the Henderson lab’s push to higher resolution. Soon, his interest was captured by single-particle cryo-EM, which could not then be used to build a three-dimensional atomic model. Two main research groups had developed software (SPIDER from Joachim Frank and IMAGIC from Marin van Heel). Grigorieff found a few processing steps that could be done more efficiently, so he began to develop what would become Frealign, a computer program to refine single-particle structures and correct for image distortions commonly found in electron microscope images.
After five years at MRC Laboratory of Molecular Biology, Grigorieff heard about a cryo-EM opening at Brandeis University near Boston, Mass. He was ready for a new adventure, and a move to the United States would put him closer to his then girlfriend in Arizona. He joined the Brandeis faculty in 1999 and one year later secured long-term funding as an investigator of the Howard Hughes Medical Institute.
“As a Hughes investigator, you can have crazy ideas,” he says. “We could try out things that otherwise would have been harder or impossible. It’s an enormous freedom.”
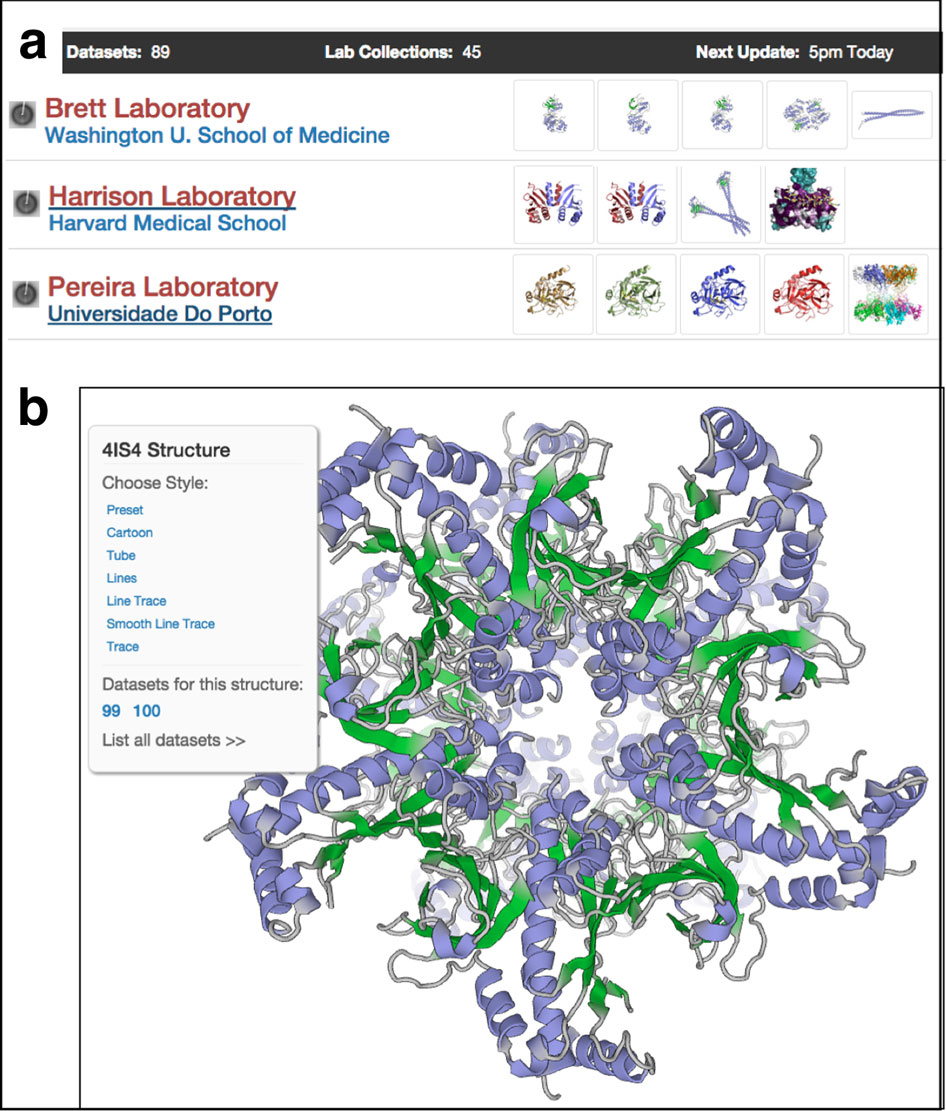
At Brandeis, he continued to develop Frealign. He also developed other single-particle EM tools and applied them in collaborations with other structural biology groups. One milestone was CTFFIND, an important step in achieving accurate high resolution with cryo-EM that became the industry standard. These tools have been refined and integrated with many others in a complete user-friendly software package called cisTEM, published in eLife March 2018, that covers the pipeline from imaging to 3D reconstruction for single particle EM.
At home, Grigorieff can sit for hours in the backyard with a telescope and a beer, looking at the quiet night sky and photographing celestial objects. Like samples caught in the cryo-EM beam, objects move across the sky. To take a photograph, a computer takes the motion into account, guiding the telescope and camera to track the object over the hours needed to collect an image.

For proteins shaken up by the electron beam, several scientists had proposed taking movies instead of pictures and aligning the movie frames to undo the blurring that normally occurs.
Suddenly, in 2011, there was a new experimental technology that could achieve this. Called a direct detector, the camera directly and quickly recorded electrons reflected off the protein sample. Three companies were developing direct detectors in collaboration with research groups, one of which was located in San Diego. Time for a sabbatical. In Boston, Grigorieff had met and married the niece of the immigration lawyer handling his green card, the children’s book author Maria Gianferrari. “We packed up the whole family, dog included, and drove from Massachusetts to San Diego for 13 months,” he says.
The sabbatical project resulted in the first report of this kind of movie collection and image correction in 2012 in the Journal of Structural Biology. First author Axel Brilot, a graduate student with Grigorieff at Brandeis, was honored with a "Paper of the Year" award in 2015. And in the scientific background for the 2017 Nobel Prize in Chemistry for development of cryo-EM awarded to Henderson, Frank, and Jacque Dubochet, the 2012 paper was cited as one of the recent advances that built on the earlier groundbreaking work by the three scientists. (The follow-up paper that demonstrated how well the technique works was rejected twice before being published.)
In 2012, Grigorieff returned to Brandeis. He realized cryo-EM was about to change, with expensive new microscopes and detectors that needed a critical mass of scientists for support. He could have tried to forge a bigger alliance in the Boston area, but he had a connection with HHMI’s Janelia Research Campus. He had helped install the first big cryo-EM instrument and had nearly accepted an offer to be a founding investigator there a decade earlier. It made more sense to him to take advantage of a new shared facility for investigators at Janelia in Virginia, where he could also devote 100% of his time to research.
With his HHMI funding recently renewed, Grigorieff will move the family back to New England in 2020, when his daughter finishes high school. This time, he will work at University of Massachusetts Medical School in Worcester. He is contemplating his next big scientific direction.
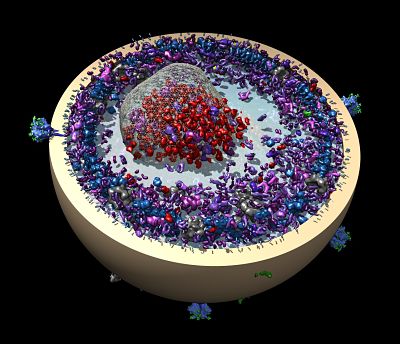
He may follow up on a feasibility study to image proteins inside cells proposed with collaborators Peter Rickgauer and Winfried Denk of the Max Planck Institute of Neurobiology in Germany. They outlined their approach in a May 2017 paper in eLife. The paper proposed a way of studying single proteins in the crowded environment of the cell, using single particle cryo-EM to catch them in interactions with other molecules.
Structural biologists purify proteins they want to study, making inferences about their function and activity from the shape and chemistry of the atomic structure. Cell biologists wonder how scientists can be sure proteins work that way inside living cells, Grigorieff says. “There’s a lot you can learn from purified protein, but it’s a valid point.”
There are many caveats and obstacles to overcome. The structure of the proteins would need to be known to find them inside the cell, they would have to be on the bigger side, and the cells themselves would be frozen, stopping all the interesting movements. Yet, it should be possible to determine what is next to what and how molecules interact in these cells to bring them to life, says Grigorieff. “It’s early days.”
--Carol Cruzan Morton