From Actin to Action
Emil Pai
University of Toronto, Canada
Published January 11, 2013
Emil Pai trained as a classical chemist in the mid-1970s at the University of Heidelberg. He spent his time learning messy, inefficiently named chemical reactions. "A 60 percent yield was cause for celebration," he says.
Then he attended a lecture on enzyme catalysis, a way to perform very precise biochemical reactions. "For a chemist, it was a humbling experience," says Pai. "I realized that to understand these reactions, you have to know what the molecules look like." Pai, now a professor of biochemistry, medical biophysics and molecular genetics at the University of Toronto (and also former PhD advisor of SBGrid director, Piotr Sliz), has been solving structures ever since.
At the Max Planck Institute for Medical Research, Pai earned his PhD, learned X-ray crystallography from Georg Schulz, and learned to make protein crystals from Heiner Schirmer. He also sat near Wolfgang Kabsch, often acting as a guinea pig for his XDS software. "Even today, I know my XDS," he says.
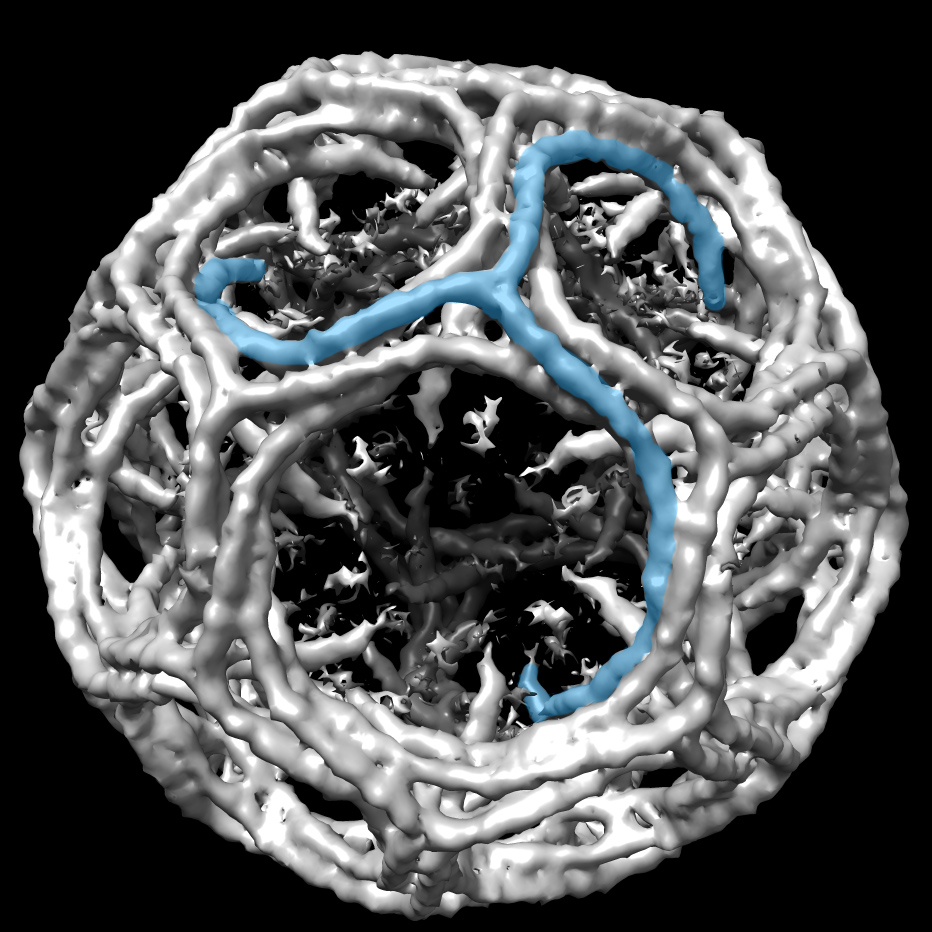
Pai and Kabsch were part of a group that collaborated to solve the first actin structure, a challenge because muscle proteins polymerize and don't form crystals. When a colleague observed that mixing actin with DNase stabilizes the protein, the group applied the technique to get a crystal. It took another 15 years to solve the structure.
In 1991, Pai moved to the University of Toronto to help build up a structural biology community there. Today there are 10 other structural biology labs spread across Toronto, all members of the SBGrid Consortium, with several investigators using NMR and electron microscopy to determine protein structures.
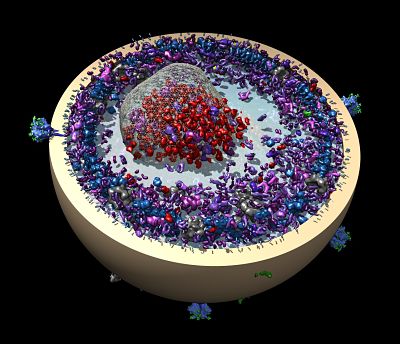
In Toronto, Pai applies crystallography to gain a more complete understanding of an HIV neutralizing antibody that may help in the design of an HIV vaccine.
He is also working to understand a magnesium channel membrane protein, corA. In late 2012, he published a paper in the Proceedings of the National Academy of Science describing a partially opened channel. Though magnesium is correlated with disease, the channel isn't a target for any medical applications yet. "You have to be cautious with magnesium," says Pai. "It's all over the place. If you remove it, you don't have a targeted change."
While still in Heidelberg, Pai had participated in applying a technique called time-resolved crystallography to p21, an oncoprotein, making it possible to watch the structure change over time, as in a video. In the past few years, he renewed his interest in this work. "We hit on a system that we thought would be perfect for time-resolved crystallography," he says.
The system is a defluorinase, an enzyme that breaks carbon-fluorine bonds, the strongest bond in organic chemistry and a barrier to cleaning up environmental pollution. So far, Pai has found that the enzyme and its complexes crystallize and diffract at high resolutions, he can make a lot of it, and he is also able to run the chemical reaction that moves the compound's parts within the crystal without destroying it.
The only thing missing is a good trigger to initiate the reaction. "Sometimes the chemistry does you in," says Pai. He is hoping to find an organic chemist up to the challenge of solving the problem.
- Elizabeth Dougherty