Supported Applications
Scipion
-
Description
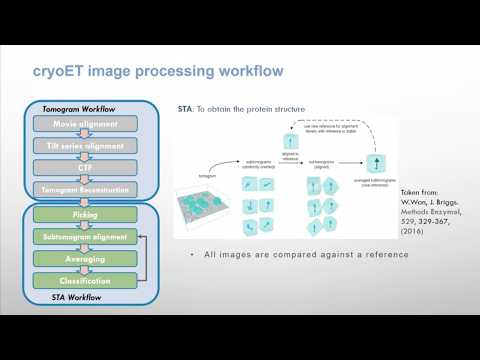
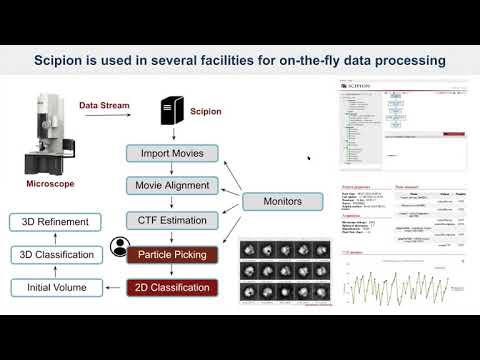
an image processing framework to obtain 3D models of macromolecular complexes using 3D EM that allows you to execute workflows combining different software tools, while taking care of formats and conversions.
-
Usage
To list all executables provided by Scipion, run:$ sbgrid-list scipion -
Usage Notes
The scipion-em-cryoSPARC2 plugin is provided in Scipion, but CryoSPARC is not available in SBGrid, so you can not use CryoSPARC directly. However, if you want to use CryoSPARC in Scipion, link a CryoSPARC installation outside of SBGrid to the scipion-em-cryoSPARC2 plugin following the instructions on its GitHub repository (skip step 1): https://github.com/scipion-em/scipion-em-cryosparc2?tab=readme-ov-file#installing-the-plugin.
SCIPION uses software that needs Nvidia GPU acceleration. Appropriate hardware and driver are required.
-
Installation
Use the following command to install this title with the CLI client:$ sbgrid-cli install scipionAvailable operating systems: Linux 64 -
Primary Citation*
J. M. de l. Rosa-Trevín, A. Quintana, L. d. Cano, A. Zaldívar, I. Foche, J. Gutiérrez, J. Gómez-Blanco, J. Burguet-Castell, J. Cuenca-Alba, V. Abrishami, J. Vargas, J. Otón, G. Sharov, J. L. Vilas, J. Navas, P. Conesa, M. Kazemi, R. Marabini, C. O. S. Sorzano, and J. M. Carazo. 2016. Scipion: A software framework toward integration, reproducibility and validation in 3D electron microscopy. Journal of Structural Biology. 195(1): 93-99.
-
*Full citation information available through
-
-
Citation Note
Scipion is freely available and its source code is 100% open. In exchange, please cite, not only us at Scipion, but ALL of the software that helped you to produce your article: http://scipion.i2pc.es/report_protocols/packages -
Webinars
SBGrid webinars are hosted with partial support from the NIH R25 Continuing Education for Structural Biology Mentors #GM151273, in collaboration with Co-PI Jamaine Davis of Belmont University.
Topic:
Image processing made simple with ScipionTomo.
Jose Luis Vilas, Ph.D.
Investigator, National Biotechnology Center, CSIC
Host: Estrella Fernandez-Gimenez
Recorded on April 14, 2026
For more information on ScipionTomo:
https://sbgrid.org/software/titles/scipionTopic: ScipionTomo: Towards cryo-electron tomography software integration, reproducibility, and validation.
Presenter: Pablo Conesa Team Leader, Spanish National Research Council
Host: Jason Key
Recorded on October 10, 2023
For more information on Scipion:
https://sbgrid.org/software/titles/scipion
https://github.com/pconesa
-
Keywords
-
Default Versions
Linux 64: 3.11.6 (42.7 GB)
-
Other Versions
Linux 64:
3.7.1 (19.3 GB)
Developers
Jose-Maria Carazo, Jose Miguel de la Rosa Trevin, Carlos Oscar Sorzano