Wire Models, Wired
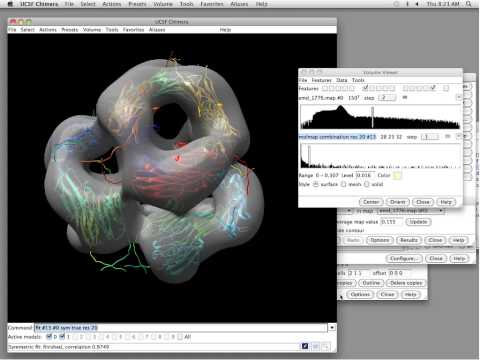
The molecular graphics software called Chimera, written and supported by a team of scientists in Tom Ferrin’s lab at the University of California, San Francisco (UCSF), has been cited over 7000 times and helps biologists and drug developers visualize molecules and biological structures in 3D at various resolutions. The tool has a personal history that traces back to 1994, and an ancestral history that stretches nearly four full decades earlier, to a London lab in 1955 and a man named Robert Langridge, also known as the pioneer of molecular graphics.

Though computation has changed almost indescribably since the middle of the 20th century, Ferrin faces some of the same challenges …