Unplanned Pioneer
Tim Stevens
Babraham Institute
Published January 15, 2013
When Tim Stevens finished his PhD in biochemistry at the University of Cambridge in 1999, he needed a job to tide him over for a few months. When he discovered that his department had 9 months of grant funding for someone to do Nuclear Magnetic Resonance Imaging (NMR) analysis, he applied.
Even though he'd never done NMR work before, he got the job, and so defined the next decade of his career.
During that 9-month stint, Stevens solved one structure on his own and assisted with another. "I'd done a lot of computing work and I knew my way around a protein very well," he says, "so using the NMR software came naturally to me."
The software Stevens was using had been written in the 1980s. "It was very good, but old fashioned," says Stevens. "I began mumbling that I could do better."
Meanwhile, on the floor just below him, Ernest Laue, a professor of structural biology at Cambridge, had a similar notion. He wanted to create a modern NMR software suite for structural biology. Patterned after CCP4 in funding and philosophy, he called it CCPN. He was looking for people to join his team. Stevens, with his computing, structural biology and new NMR experience, was a perfect fit.
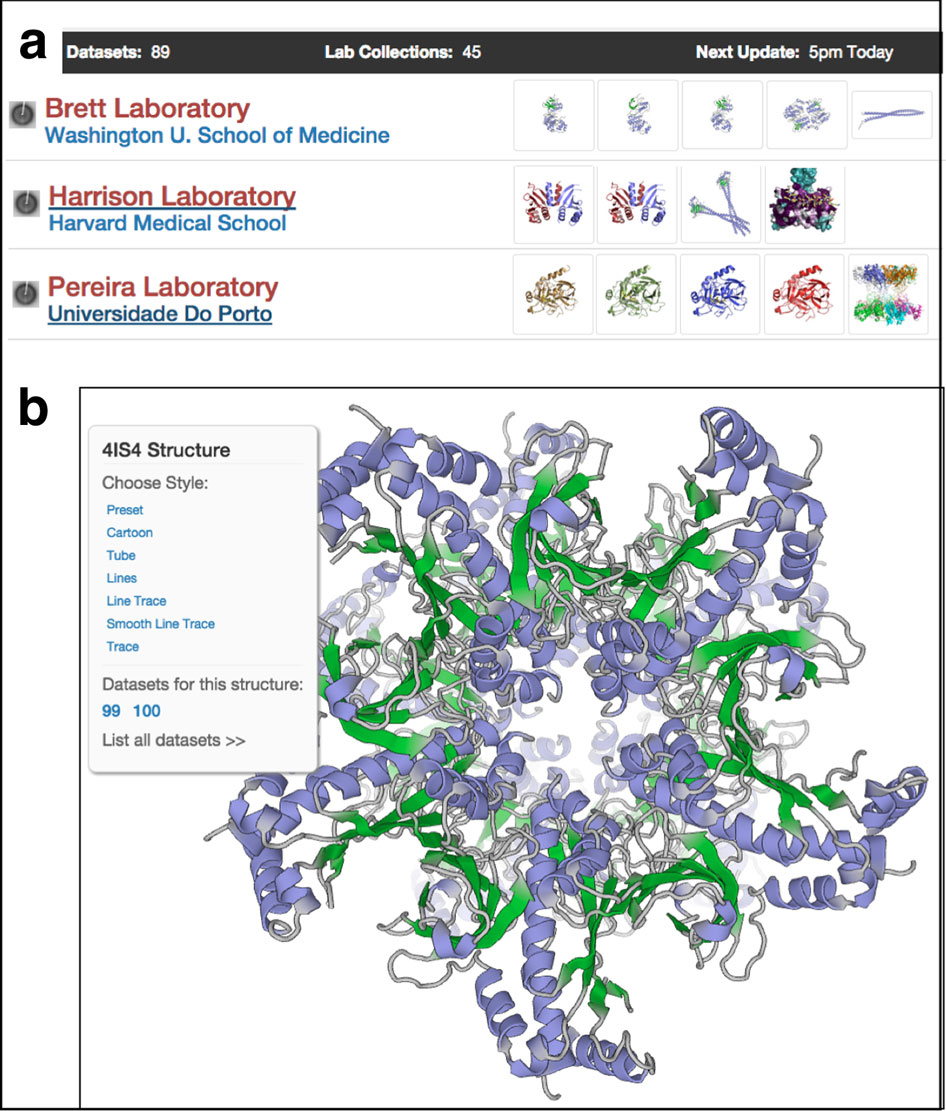
Stevens joined Rasmus Fogh and Wayne Boucher, two other postdoctoral fellows working on the new software suite. Stevens focused on applications, such as the Analysis graphical tool. Within a few years, they had a product.
Today, CCPN has about 1000 users. "We've had great interactions with the user community," says Stevens, who has traveled around the world to participate in CCPN workshops.
Recently, Stevens has focused on retooling the user interface, taking it through yet another round of modernization. "That work is about halfway done," he says. He debuted SpecView and ChemBuild, previews of CCPN version 3, in an SBGrid Webinar in 2012.
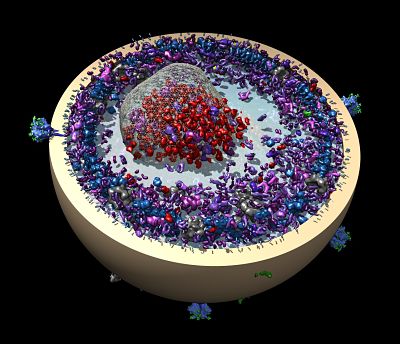
Stevens is quick to point out that computing isn't his only gig. "I'm a bioinformatician that dabbles in NMR analysis," he says. During the past decade, Stevens has continued his work characterizing membrane proteins and their distinguishing features.
He has also worked on applying NMR to metabolomics. Unlike Xray crystallography, which essentially snaps a picture of a molecule, NMR spectra contain a jumble of signals that must be pieced together like a jigsaw puzzle. Those signals can also be useful indicators for screening, such as scanning blood or urine for specific compounds or scanning proteins to detect strong or weak binding with a drug candidate. "NMR is very sensitive and can detect even subtle changes. You can even measure protein folding," he says. "You can do an awful lot with it."
Stevens left his long-time post at CCPN at the end of 2012 due to funding constraints. The change is somewhat fortuitous, allowing him to transition to a position at the Babraham Institute, potentially defining the coming decade of his career.
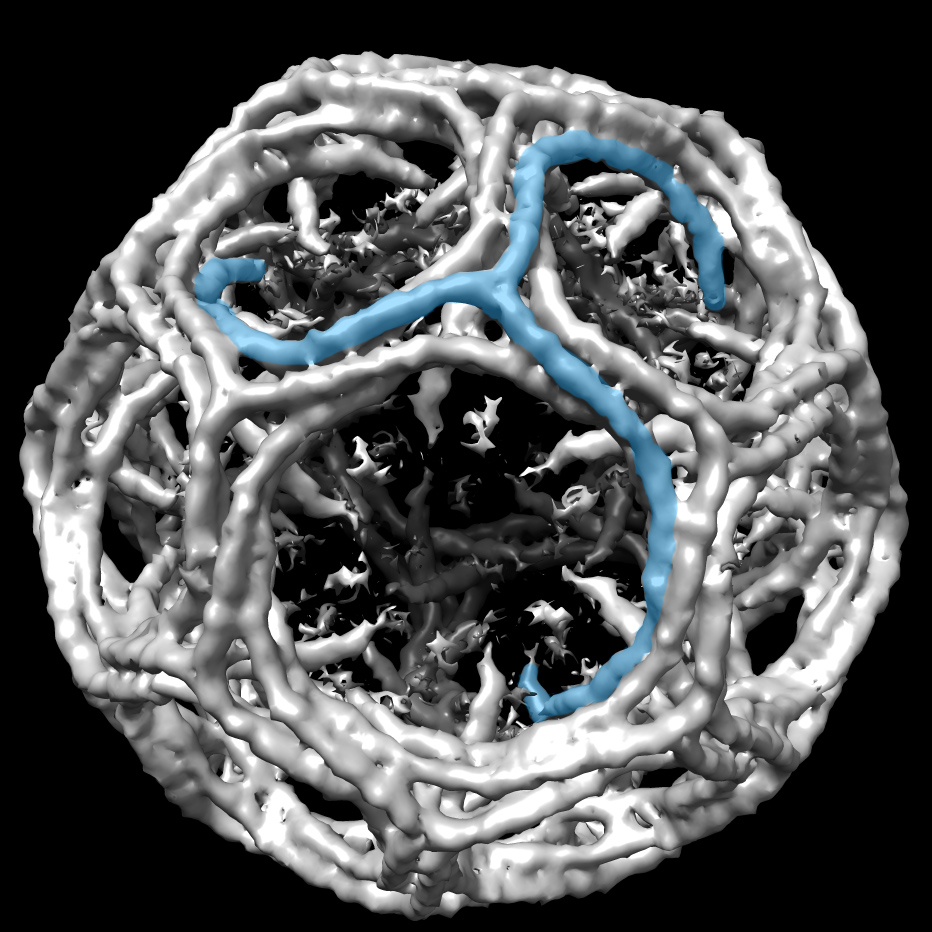
At Babraham, Stevens will apply the analysis techniques of NMR to look at the macro structure of chromosomes. He and his colleagues hope to determine whether physical locations of genes contribute to their regulation. "When DNA is damaged, does that affect the physical arrangements of genes? If so, does that contribute to cancer or aging?" he asks.
To answer these questions, Stevens will combine his expertise in genomic sequencing and bioinformatics, structural biology and NMR analysis. "It's somewhat new territory," he says. "But we're hitting the technology at just the right stage."
- Elizabeth Dougherty